In the few weeks since we announced the Lance-Hugging Face Hub integration, we’ve seen numerous examples of community members uploading their own Lance datasets to the 🤗 Hub. See this page for some recent examples. 🚀
If you’re looking to upload your own Lance datasets to Hugging Face but aren’t sure how, this post is for you. We’ll describe, step-by-step, how to create, query, manage, and share your open-source datasets in Lance format with the world, and why that matters. Let’s dive in.
Example Scenario
Say you have a dataset of magical characters from the kingdom of Camelot. The dataset is multimodal, consisting of JSON blobs with each character’s metadata and their images (JPG files). If you’ve never read up on Arthurian legend, here are the images and names of some of the important characters:
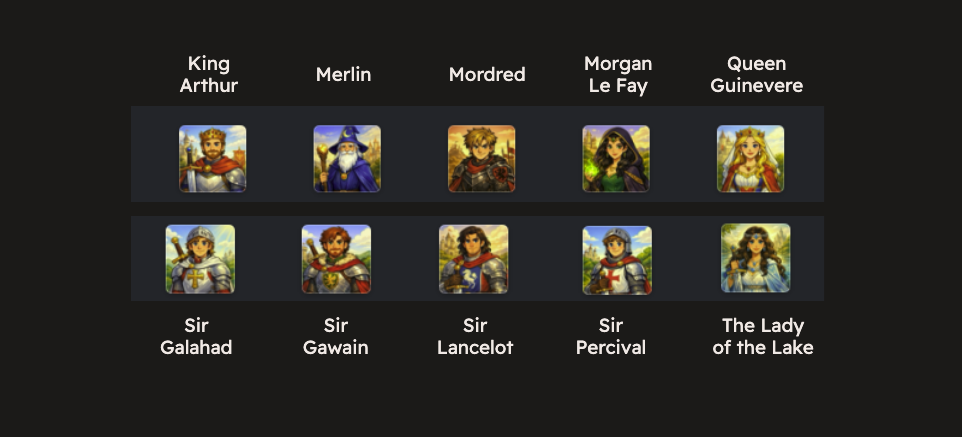
We see familiar characters like Merlin, Arthur, his wife Queen Guinevere (also known as Geneva), and Sir Lancelot 🗡️. The metadata for each character is available as JSON (see the full file here ).
[
{
"id": 1,
"name": "King Arthur",
"role": "King of Camelot",
"description": "The legendary ruler of Camelot, wielder of Excalibur, and leader of the Knights of the Round Table.",
"stats": {
"strength": 2,
"courage": 5,
"magic": 1,
"wisdom": 4
}
},
{
"id": 2,
"name": "Merlin",
"role": "Wizard and Advisor",
"description": "A powerful wizard and prophet who mentors Arthur and shapes the destiny of Camelot through magic and foresight.",
"stats": {
"strength": 2,
"courage": 4,
"magic": 5,
"wisdom": 5
}
},
// ... more character info below
]The Problem
The simplest way to share this dataset with others on the Hugging Face Hub would be to just upload the raw files (JSON + images in a separate directory) and be done with it. However, this has the following problems:
- For users to explore the metadata using filters, keyword-based search and vector search, they need to download the full dataset locally and write ad-hoc scripts to isolate parts that are useful.
- Images and other multimodal assets (video, audio, PDFs) contain rich information, and they aren’t natively “searchable” unless you already have embeddings of them.
For large datasets that are several terabytes in size, it’s easy to imagine how much repetitive (and expensive) work is required to consume a naively uploaded dataset. In many domains like genomics, robotics, and autonomous vehicles, you routinely see billions of records with rich information in nested fields, images, and video files, so it’s important to consider how the data is stored and packaged for downstream tasks like model training or search.
Below, we demonstrate how storing your data in Lance format on the Hub solves these problems.
Why Lance?
Lance is an open-source, columnar file and table format that makes querying, managing, and distributing AI datasets simple. When you store your data in Lance format, you get the following benefits:
- Store multimodal assets as first-class citizens, managed as part of the dataset lifecycle.
- Store indexes (vector, full-text search and scalar indexes) as part of the dataset.
- Zero-copy data evolution, meaning you can easily add derived columns (like features or embeddings) at a later time, without full table rewrites. Only data that is new, is written – while expensive existing data (like images/videos) remain untouched.
- Built-in automatic data versioning with time-travel across versions
LanceDB is a multimodal lakehouse and an embedded retrieval library that offers a convenient interface on top of the Lance format for ML/AI engineers, for numerous tasks ranging from training, feature engineering, search and analytics.
By sharing your AI datasets in Lance format on the Hugging Face Hub, users can instantly scan, filter, and search the data via LanceDB, without having to first download the dataset locally.
The upcoming sections will illustrate how to do each of these steps in combination with the Hugging Face CLI.
Walkthrough
In this section, we’ll demonstrate how to build a multimodal Lance dataset with vector and FTS indexes for our Camelot characters, publish the full dataset (including indexes) to the Hugging Face Hub, and scan/query it directly from the Hub using LanceDB. If you want to follow along, see the code to reproduce the below workflow here .
We’ll be using an OpenAI text embedding model in this workflow, so we’ll export an OPENAI_API_KEY. Although not explicitly required for this small dataset, rate limits can be an issue when working with larger datasets, so we also recommend exporting an HF_TOKEN and authorizing it via the CLI as follows:
export OPENAI_API_KEY=your_openai_key_here
export HF_TOKEN=your_hf_write_token_here
hf auth login --token $HF_TOKENStep 1: Transform raw data into Lance tables
The first step is to design a table schema in LanceDB that captures the necessary fields for downstream use cases. The data model shown here uses just one Lance table that contains all the data, including image blobs for each Camelot character.
LanceDB (and the Lance format) are Arrow-native, so we define a PyArrow schema as follows:
import pyarrow as pa
schema = pa.schema(
[
pa.field("id", pa.int32(), nullable=False),
pa.field("image", pa.binary(), nullable=False),
pa.field("name", pa.string(), nullable=False),
pa.field("role", pa.string(), nullable=False),
pa.field("description", pa.string(), nullable=False),
pa.field(
"stats",
pa.struct(
[
pa.field("strength", pa.int8()),
pa.field("courage", pa.int8()),
pa.field("magic", pa.int8()),
pa.field("wisdom", pa.int8()),
]
),
nullable=False,
),
pa.field("image_path", pa.string(), nullable=False),
pa.field("image_vector", pa.list_(pa.float32(), list_size=IMAGE_DIMS)),
pa.field("text_for_embedding", pa.string(), nullable=False),
pa.field("text_vector", pa.list_(pa.float32(), list_size=TEXT_DIMS)),
]
)The Arrow schema shown above mimics the JSON metadata structure, including nested fields (stats) that are stored as structs in PyArrow and can be queried directly in LanceDB without needing to unnest them.
We also add the following new columns as part of the schema:
image: Image bytes, of thebinaryArrow typeimage_vector: Image embedding, generated using the OpenCLIP embedding modeltext_vector: Text embedding on thedescriptionvalues, generated using the OpenAItext-embedding-3-smallmodel
Note that the image bytes are binary representations (blobs) that are stored inline and versioned with the rest of the data – because multimodal data is a first-class citizen in Lance. The two vector columns are derived columns, from the image and text_for_embedding columns, respectively.
Tip: Use PyArrow RecordBatches
It’s strongly recommended to ingest data in batches of PyArrow records via iterators rather than looping through each item row by row.
def iter_row_batches(batch_size: int = 5) -> Iterator[pa.RecordBatch]:
# Gather source data as a list of dicts
records = sorted(json.loads(INPUT_JSON.read_text()), key=lambda r: r["name"])
image_paths = sorted(IMAGE_DIR.glob("*.jpg"), key=lambda p: p.name)
source_rows: list[dict] = []
for row, img_path in zip(records, image_paths):
stats = row["stats"]
source_rows.append(
{
"id": row["id"],
"name": row["name"],
"role": row["role"],
"description": row["description"],
"stats": {
"strength": stats["strength"],
"courage": stats["courage"],
"magic": stats["magic"],
"wisdom": stats["wisdom"],
},
"image_path": f"raw_data/img/{img_path.name}",
"image": img_path.read_bytes(),
"text_for_embedding": f"{row['role']}. {row['description']}",
}
)
# Yield PyArrow record batches as iterators
for start in range(0, len(source_rows), batch_size):
chunk = source_rows[start : start + batch_size]
text_vectors = embed_texts([row["text_for_embedding"] for row in chunk])
image_vectors = embed_images([row["image"] for row in chunk])
out_rows = []
for row, txt_vec, img_vec in zip(chunk, text_vectors, image_vectors):
out_rows.append(
{
**row,
"text_vector": txt_vec,
"image_vector": img_vec,
}
)
yield pa.RecordBatch.from_pylist(out_rows, schema=schema)The batch size parameter determines how large each batch is, prior to ingestion. It’s set to be 5 here to demonstrate more than one batch being ingested, but in practice, you would set a much larger value (10K or more, depending on the data).
When creating the Lance dataset, use the create_table method on the LanceDB connection.
For the data parameter, instead of passing a fully materialized data object, pass in the iter_row_batches() function. This yields PyArrow records in batches during ingestion.
import lancedb
db = lancedb.connect("./magical_kingdom")
table = db.create_table(
TABLE_NAME,
data=iter_row_batches(),
schema=schema,
mode="create", # "overwrite" if you want to start from a clean slate
)Every write operation (via a transaction) in Lance/LanceDB creates a new version of the dataset. By yielding the data via iterators, LanceDB lazily consumes batches of the data without materializing it in full, and only one new version of the dataset is created after the script finishes.
In contrast, naively looping through each record and inserting new rows via table.add
will create a new version for each write – if inserting thousands of records at a time, this will result in thousands of versions of the dataset, which isn’t ideal from a data management perspective. Versions should be representative of macro-operations on the dataset, so it’s worth creating versions each time a workflow is run (rather than each time a new row is inserted).
Once create_table finishes running, you can also create indexes. In this project, we’ll create an FTS index on the description column:
table.create_fts_index("description", replace=True)For this small a dataset, no vector index is necessary, but for real-world datasets that are tens of thousands of rows or more, you can create a vector index on the required vector columns as shown here .
We now have our local Lance dataset (metadata, image bytes, indexes and versions) in a table named characters.lance. A single Lance dataset packages the data files, indexes, and transaction and version metadata all in one place.
magical_kingdom/
└── characters.lance/
├── _indices/
├── _transactions/
├── _versions/
└── data/Step 2: Upload Lance dataset to Hugging Face Hub
The next step is to upload the Lance dataset to Hugging Face. We’ll use the
Hugging Face CLI
command below from the root directory. Ensure that you authenticate your CLI with a valid HF_TOKEN with write access.
The Hugging Face CLI comes with a upload-large-folder flag which uses a resumable multi-commit flow, making it more flexible and error-tolerant than the default hf upload:
# Upload a "dataset" repo with the name `magical_kingdom` to the Hub
hf upload-large-folder lancedb/magical_kingdom magical_kingdom \
--repo-type dataset \
--revision mainThis creates a new dataset repo on the Hub where 9 files were hashed and uploaded.
Files: hashed 1/9 (1.1K/2.7M) | pre-uploaded: 0/0 (0.0/2.7M) (+9 unsure) | committed: 0/9 (0.0/2.7M) | ignored: 0
Files: hashed 9/9 (2.7M/2.7M) | pre-uploaded: 1/1 (2.7M/2.7M) | committed: 9/9 (2.7M/2.7M) | ignored: 0Next, you can inspect the existing versions of the dataset on the Hub. The following script does this:
from pprint import pprint
import lancedb
db = lancedb.connect("hf://datasets/lancedb/magical_kingdom")
table = db.open_table("characters")
versions = table.list_versions()
pprint(versions)This returns:
[{'metadata': {'total_data_file_rows': '10',
'total_data_files': '1',
'total_deletion_file_rows': '0',
'total_deletion_files': '0',
'total_files_size': '2663804',
'total_fragments': '1',
'total_rows': '10'},
'timestamp': datetime.datetime(2026, 3, 5, 9, 52, 52, 173784),
'version': 1},
{'metadata': {'total_data_file_rows': '10',
'total_data_files': '1',
'total_deletion_file_rows': '0',
'total_deletion_files': '0',
'total_files_size': '2663804',
'total_fragments': '1',
'total_rows': '10'},
'timestamp': datetime.datetime(2026, 3, 5, 9, 52, 52, 185318),
'version': 2}]Each commit to a Lance table via a transaction creates a new version of the dataset. The first version is created when we run create_table and the data is ingested. The second version is created when we create the FTS index. Any subsequent writes, updates or deletes (or index creations) will create more new versions.
Step 3: Add a new column to the dataset
In the real world, datasets are rarely static – they evolve. Imagine a scenario where you want to add a new category column to the characters table then backfill its values into the table.
This operation is both a schema update and a data update, which Lance handles efficiently. Because Lance supports incremental data evolution , it can add, remove, and alter columns without rewriting existing data files, making updates to large tables very I/O-efficient.
Let’s update the local dataset with the new column.
import lancedb
db = lancedb.connect("magical_kingdom")
table = db.open_table("characters")
# Step 1: add the new column (schema evolution).
table.add_columns(pa.field("category", pa.string()))This updates the schema of the Lance dataset, creating a new version.
For updating the dataset, we use a classify function based on the role and description fields to assign a character a category.
Only one of five categories is allowed: “king”, “queen”, “knight”, “mage” and “other”.
# Step 2: Merge-insert the category values to the Lance table
def classify(role: str, description: str) -> str:
"""Assign a category from role + description using simple heuristics."""
role_l = role.lower()
text_l = f"{role} {description}".lower()
if "king" in role_l:
return "king"
if "queen" in role_l:
return "queen"
if "knight" in role_l:
return "knight"
if any(token in text_l for token in ["wizard", "sorcer", "mage", "enchant", "magic"]):
return "mage"
return "other"
# Create a new PyArrow table with "id" as the join key
category_data = pa.table(
{
"id": source.column("id"),
"category": pa.array(categories, type=pa.string()),
}
)
# Step 3: one merge_insert to update category by id.
(
table.merge_insert("id")
.when_matched_update_all()
.execute(category_data)
)Using merge_insert lets you specify id as the join key while passing in a precomputed list of string values for the new category column. In this example, we use simple hardcoded rules, but in a real-world scenario, it’s common to use ML models or LLMs to compute derived column values from existing ones.
LanceDB Enterprise users can use
Geneva
, a multimodal feature engineering toolbox that provides versioned UDFs that run in parallel, via Ray, either locally on your laptop or on an Enterprise LanceDB cluster. With Geneva, feature engineering becomes as simple as decorating your existing function and calling table.backfill to add derived columns. You can run a pipeline on a few records as easily as you run it on billions of rows, with just a few lines of code.
@udf
def classify(role: str, description: str) -> str:
# Rules or ML model to classify a character here ...
return category
# Register computed column from the UDF
tbl.add_columns({"category": classify})
# Run distributed backfill for that computed column
tbl.backfill("category")Step 4: Upload the modified dataset
Now that the new column and its values are added to the dataset, upload it back to the Hub with a new commit message:
hf upload-large-folder lancedb/magical_kingdom magical_kingdom \
--repo-type dataset \
--revision mainWhen this dataset is uploaded, even though we included the entire directory, as the log below shows, 5 new files are detected. Hugging Face’s storage backend (in combination with Lance’s efficient data evolution) avoids wasted I/O by only uploading what’s changed (i.e., the 9 files that existed before aren’t re-uploaded).
Files: hashed 9/14 (2.7M/2.7M) | pre-uploaded: 1/1 (2.7M/2.7M) (+5 unsure) | committed: 9/14 (2.7M/2.7M) | ignored: 0
Files: hashed 14/14 (2.7M/2.7M) | pre-uploaded: 1/1 (2.7M/2.7M) | committed: 14/14 (2.7M/2.7M) | ignored: 0The version history now shows that two new versions of the dataset are present (versions 3 and 4). Version 3 is for when we added the new column (schema update), and version 4 is for when the new values were added (data update).
[{'metadata': {'total_data_file_rows': '10',
'total_data_files': '1',
'total_deletion_file_rows': '0',
'total_deletion_files': '0',
'total_files_size': '2663804',
'total_fragments': '1',
'total_rows': '10'},
'timestamp': datetime.datetime(2026, 3, 5, 9, 53, 15, 360538),
'version': 3},
{'metadata': {'total_data_file_rows': '10',
'total_data_files': '2',
'total_deletion_file_rows': '0',
'total_deletion_files': '0',
'total_files_size': '2664444',
'total_fragments': '1',
'total_rows': '10'},
'timestamp': datetime.datetime(2026, 3, 5, 9, 53, 15, 365071),
'version': 4}]Manage dataset versions
You can time-travel between versions of datasets in Lance with table.checkout. The example below shows how to check out the first version
of the dataset.
# Read the first version of the table
first_version = min(v["version"] for v in versions) # oldest/first manifest version
table.checkout(first_version)
print(table.version) # now reading old versionSimilarly, you can restore past versions of the dataset as the current version:
# Restore the first version as the latest version
# Creates a new latest version equal to first_version's data
table.restore(first_version) # First version is now the latest versionOver time, as you accumulate more and more versions, you can run a
compaction
job by running the command table.optimize() – this cleans up old versions and keeps your dataset size reasonable.
Query the dataset on the Hub
This is the fun part! LanceDB accepts the hf:// path specifier, so you can directly scan data from the Hub, without needing to download it locally.
The example below shows how to open the characters table on the Hub via the remote path and run a query with filters for the strongest knight. Note that strength is a nested field under stats (which is a struct) – LanceDB supports directly querying on nested fields and structs – simply pass it in as an object in the query (e.g., stats.strength).
import lancedb
# Scan data directly from the Hugging Face Hub
# (No need to download the dataset locally)
db = lancedb.connect("hf://datasets/lancedb/magical_kingdom")
table = db.open_table("characters")
r = table.search() \
.where("category = 'knight'") \
.select(["name", "role", "stats.strength"]) \
.limit(5) \
.to_polars() \
.sort("stats.strength", descending=True) \
.head(1)The knight with the greatest strength in Camelot is Sir Lancelot! 🗡️
| name | role | stats.strength | image |
|---|---|---|---|
| Sir Lancelot | Knight of the Round Table | 5 |  |
Next, let’s do a full-text search for the keyword “mysterious”.
keyword = "mysterious"
return (
table.search(keyword, query_type="fts")
.select(["name", "role", "description"])
.limit(5)
.to_polars()
)We get the Lady of the Lake as a result, who is “a mysterious supernatural figure associated with Avalon, known for giving Arthur the sword Excalibur.”
| name | role | description | image |
|---|---|---|---|
| The Lady of the Lake | Mystical Guardian | "A mysterious supernatural ..." |
 |
Finally, let’s run an image vector search for the following query:
# Define image embedding function here (OpenCLIP)
query = "a powerful mage with a staff and a long beard"
query_vector = image_embedding.compute_query_embeddings(query)[0]
(
table.search(query_vector, vector_column_name="image_vector")
.select(["name", "role", "description"])
.limit(1)
.to_polars()
)| name | role | description | image |
|---|---|---|---|
| Merlin | Wizard and Advisor | "A powerful wizard and prophet ..." |
 |
Add a dataset card
If you’re the dataset’s maintainer, it’s a good practice to include an informative dataset card once you upload it to the Hub. This communicates the schema and usage of the dataset to other developers using the dataset. The card lives at the repo’s root in a file named README.md on the Hub.
This project keeps the source card text in
HF_DATASET_CARD.md
, so you can publish updates
to the dataset there and upload it as README.md using the following command on the HF CLI:
hf upload lancedb/magical_kingdom HF_DATASET_CARD.md README.md \
--repo-type dataset \
--commit-message "Update dataset card"When you visit the dataset repo on Hugging Face , the card will immediately be visible on the home page.
Uploading your own Lance dataset
If you already have a dataset on the Hub in another format (like CSV or Parquet), now is a good time to publish a Lance version of it!
- Convert your source data into one or more Lance tables (including derived columns and indexes you want downstream users to query).
- Upload the dataset repo with
hf upload ... --repo-type dataset. - Add a clear
README.mddataset card so others can understand schema, query examples, and intended use.
Once it’s live, you can test it from a clean environment with lancedb.connect("hf://datasets/<org>/<dataset>") to confirm users can query it remotely without needing to download it locally.
Benefits for Hugging Face Users
As a Hugging Face user, the examples above highlight how convenient it is to run a variety of exploratory (and analytical) queries on a remote Lance dataset to understand which subsets of the data are important/useful for training, analytics or search use cases downstream.
Once you do, it’s trivial to download the entire dataset via the CLI and begin using it locally:
hf download lancedb/magical_kingdomThe same approach works for really massive datasets (see fineweb-edu , which contains 1.5 billion rows plus an FTS index).
Conclusions
LanceDB makes Hugging Face dataset publishing much more practical at scale for multimodal workloads: it stores and versions metadata, blobs and embeddings as part of the dataset lifecycle and supports scalable indexing and schema evolution. You can conveniently scan and query the dataset directly from the Hub, without needing to download it locally. See the full code to reproduce the above workflow here .
As you go about publishing your own datasets, keep these best practices in mind:
table.addhelps add new row values (appends/inserts) andtable.merge_inserthelps add new column values (upserts). Use either of them to modify your dataset as your needs evolve. See the LanceDB docs for more examples.- Avoid writing data with naive loops that insert or update one row at a time. Each transaction in LanceDB creates a new dataset version – yielding batches of PyArrow records and inserting/updating them in bulk is a very efficient approach that also does not unnecessarily create new versions.
- Manage dataset versions by inspecting them regularly with
table.list_versions(), especially after ingestion, schema updates, and backfills. - Keep your dataset size manageable by periodically running
table.optimize()– this compacts old fragments and trims the disk space used.
As a next step, browse through some existing Lance datasets on the Hub, publish your own, share it with the community and call us out on social media so that we can spread the word! Have fun creating your own Lance datasets on Hugging Face! 🤗




